Welcome to SuperFly!
What is SuperFly?
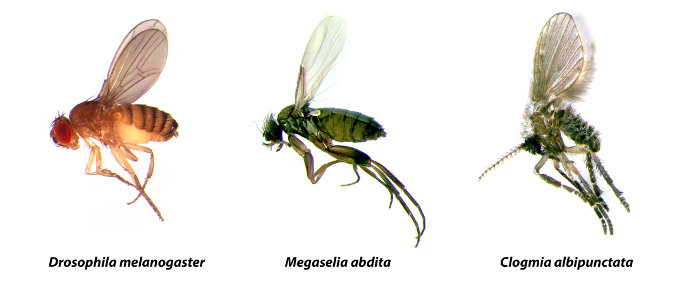
SuperFly is a database for the comparative analysis of segmentation gene expression and regulation in dipteran species (flies, midges, and mosquitoes). Currently, we provide semi-quantitative, spatio-temporal expression data for maternal co-ordinate, and gap genes in three species: the vinegar fly Drosophila melanogaster (Drosophilidae), the scuttle fly Megaselia abdita (Phoridae), and the moth midge Clogmia albipunctata (Psychodidae).
The data range from raw microscopy images of whole-mount in situ hybridisation experiments to the intermediate results of processing and quantifying the raw images, to the final integrated spatio-temporal gene expression profiles for each gene.
How to use SuperFly:
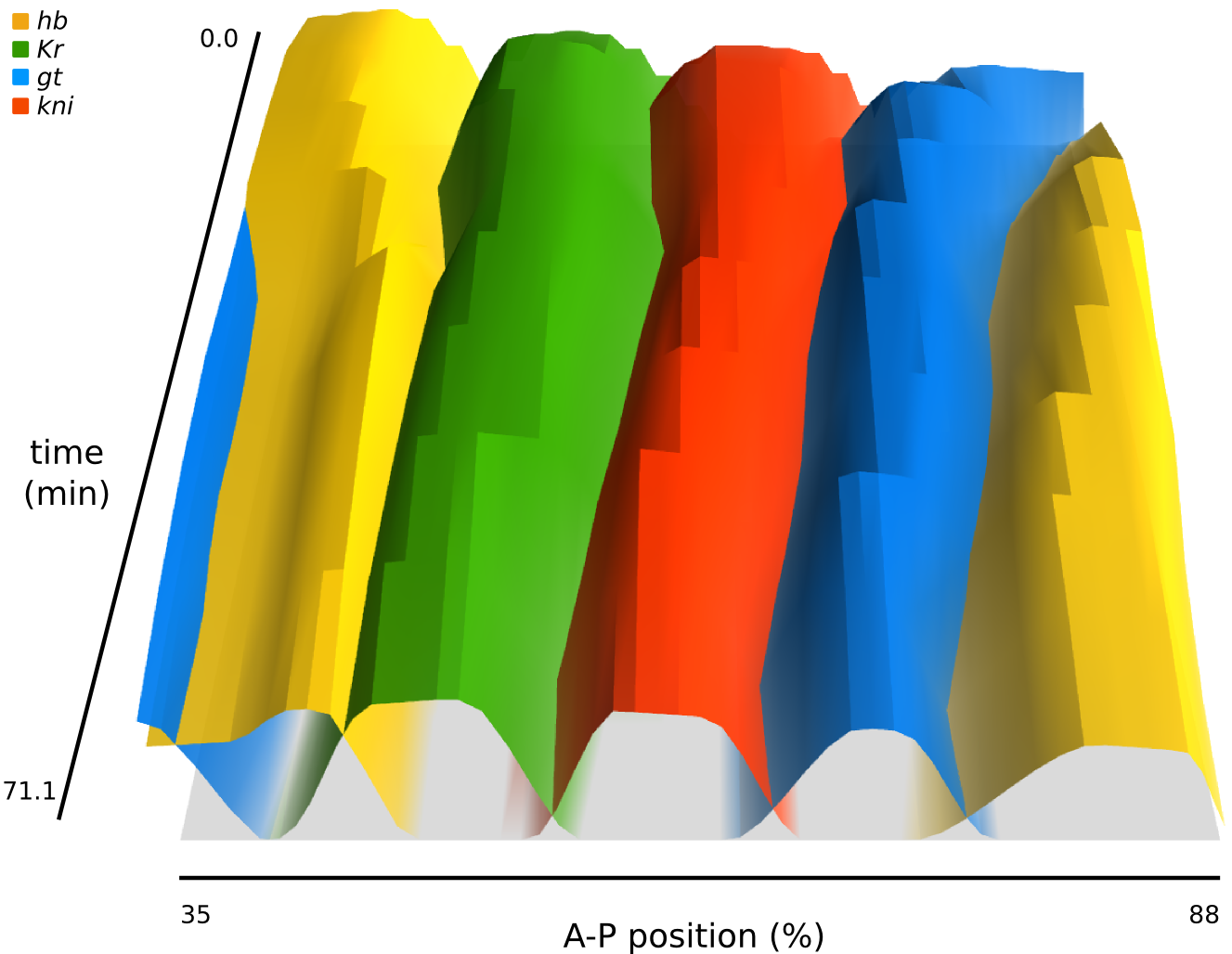
The use of SuperFly is simple and intuitive. The home page contains three main tabs. Currently, we are in the “Embryo Viewer”, as indicated in the top bar. On the left, you can select the species, gene(s), and time classes that you are interested in. Extra options are normally hidden, but can be brought into view with a single click (if you do not understand those, you should probably not worry about them!). Once you have made your selection, you can search for any embryos that match your request. The resulting list of embryos is displayed, and in case you want further information on a specific embryo, you can click “Details...” for details. The “Boundaries” tab works along similar lines, with the additional option to choose which boundary you want to view (for instance, the posterior boundary of the abdominal kni domain, etc.). Finally, the “Integrated Datasets” tab provides 2D and 3D plots of the integrated, quantified expression data (per gene), and a simple tabular format to download the data.
Details on how the data were processed can be found in the publications listed in About section.